An explanation of Estimated Breeding Values (EBVs) by Dr Tom Lewis of the Kennel Club
Analysis of genetic disease
Some genetic diseases are caused by a defective (mutant) variant (allele) at a single gene. In most cases, because genes are inherited in pairs (one from either parent), a single normal allele is sufficient to allow normal function (i.e. the mutant is recessive to the normal allele). However, when both alleles are mutants, the protein produced does not result in normal function, leading to disease. Examples of such diseases include cystic fibrosis in humans and primary lens luxation, hereditary cataract and progressive retinal atrophy in dogs. Because the disease is caused entirely by a single gene, the disease is inherited in a very predictable way in families, and it is therefore relatively easy to find the gene, identify the mutant allele and offer a DNA test.
However, most traits and diseases are a mix of multiple genes and environmental factors (for example – there is no single gene for height or blood pressure). Examples include schizophrenia in humans and hip dysplasia in dogs. The influence of multiple genes and environmental effects on the severity of disease means the disease is often less easily categorised as ‘clear’ and ‘affected’ and makes it more difficult to track within families. Selection against the disease can be slow in such cases, since we are selecting on appearance (phenotype) which is only partly determined by genetics (which is the only heritable component). Thus, phenotype is not necessarily a reliable indicator of genetics.
Statistical techniques are available which, by using pedigree information to determine relationships between dogs, discount the non–genetic/environmental influences on the disease. Also by using phenotypic information on relatives we are better able to determine an individual’s genetic risk (for example, we might re-consider using a dog with a high hip score for breeding if we knew that it already had 20 progeny all with hip scores of zero). These analyses determine the extent of genetic variation in the breed or population, and how much of the phenotypic variation this comprises. This is important in informing of how successful selection will be.
What are EBVs?
EBV stands for Estimated Breeding Value, and is the estimate of genetic risk for each animal in the pedigree from the analyses described above. It is necessary to think in terms of genetic risk because there are multiple genes each influencing the overall risk to a smaller degree. Because the EBV is the estimate of genetics alone, it is a more accurate metric for selection than phenotype (which comprises non-heritable environmental effects). More accurate selection results in a better response to selection. Furthermore, EBVs are available for all animals in the pedigree used in the analyses described above, even if they have no phenotype available.
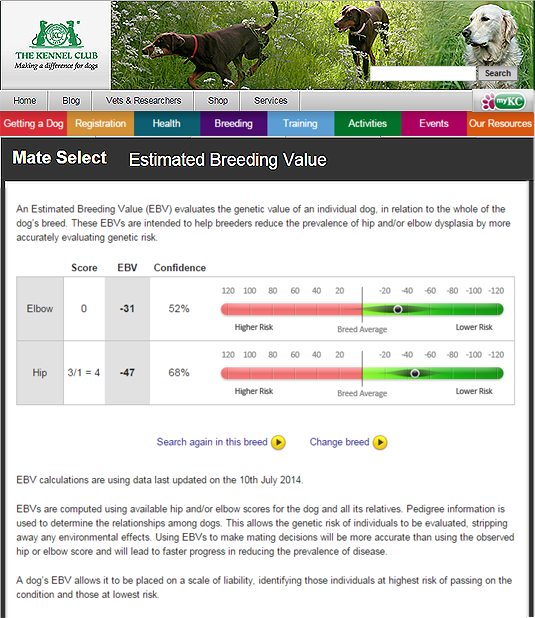
Improving the accuracy of EBVs
The EBV is an estimate of genetic risk of an individual based on the phenotypic information on itself and its relatives at the time of calculation. More information will mean we have more confidence that the estimate reflects the true value. This is computed as the accuracy. For example, an EBV for a dog which has no phenotype (e.g. hip score) for itself or any close relatives will have a lower accuracy than a dog which has phenotypic information on itself, its parents and several progeny. The best way to improve accuracy is therefore to generate as much phenotypic information as possible. The accuracy is displayed as ‘confidence’ in the above figure.
Can EBVs change?
Yes. Although the genes are fixed at conception and do not change during the lifetime of an individual, the EBV is an estimate of genetic risk, and this estimate will change as more information becomes available. Therefore, an EBV of a neonatal dog will very probably change as its own phenotype is obtained, perhaps phenotypes of siblings, and eventually phenotypes of progeny. All this extra information will increase the confidence of the estimate and may result in changes in EBV.
EBVs will also change in another way too. EBVs are usually given as a deviation from the breed or population mean. However, the average of the breed will change as the breed responds to selection over the generations. Thus while a dog’s own estimate of genetic risk may not change significantly over the course of its life, that of the wider breed may have done so in response to selection.
Considering multiple traits
Many breeds are faced with multiple heritable diseases which breeders wish to select against. While breeders may use EBVs for different diseases in isolation of each other, there are techniques available to further improve the accuracy of selection. It may be that different heritable conditions are influenced by the same genes to some degree (for example hip and elbow dysplasia in some breeds). This is important, since selection against one disease may reduce or increase the genetic risk of another. We can evaluate the extent of the genetic relationship between traits with the genetic correlation.
However, with sufficient data we can go further and use a selection index to derive ‘weights’ to be applied to EBVs for multiple traits to yield the optimal accuracy of selection for pre-defined selection objectives.
EBVs for diseases prevalent in Cavaliers
Mitral Valve Disease (MVD) and Syringomyelia are both diseases prevalent in the Cavalier, which have been shown to be heritable and so could benefit from the availability of EBVs to assist selection. Essential to allow the regular calculation of EBVs is a large quantity and regular flow of good quality phenotypic data. To date this has been accomplished via screening schemes run by the British Veterinary Association (BVA) and Kennel Club (KC; e.g. the hip dysplasia scoring scheme www.bva.co.uk/Canine-Health-Schemes/Hip-scheme). There is already an established BVA/KC Syringomyelia screening scheme (www.bva.co.uk/Canine-Health-Schemes/CM-SM-scheme), which seeks to evaluate the severity of Chiari Malformation and Syringomyelia and provide data for the calculation of EBVs. Although to date participation has been slow, every participant contributes to making EBVs for this condition more achievable.
Screening schemes for MVD in Cavaliers do exist but tend to be administered independently by individual cardiologists working with breed clubs, with no centralised data collection. This means it is difficult to ensure consistent protocols and consistent data submission. It is hoped that a BVA/KC heart screening scheme may help solve these problems.
It should be said that EBVs which cater for ‘multiple traits’, in the Cavalier breed, ie CMSM and MVD are a long way off.
